Research
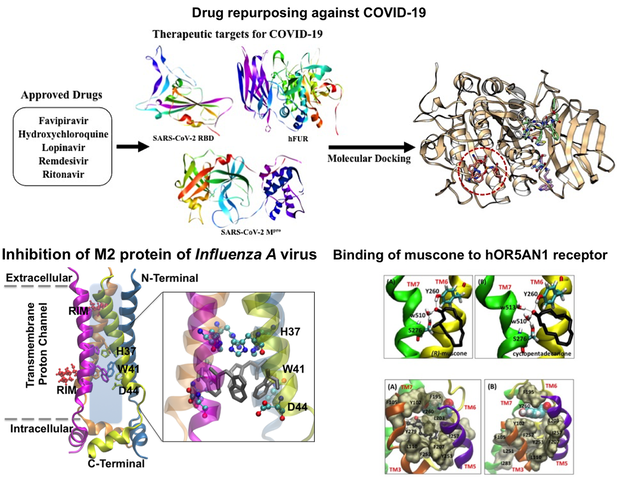
Investigation of enzyme and protein dynamics and design of small molecule modulators: We intensively focus on the 3-D structures of proteins by utilizing classical and the state-of-art constant pH molecular dynamics (MD) simulations, hybrid quantum-molecular mechanical (QMMM) calculations. Atomistic level understanding provided by these simulations/calculations reveal precious information on the modulator binding sites, hotspots on the protein and the structural outcome of the modulator binding. These data in conjunction with the experimental data form the building blocks of very important research area: drug designing.
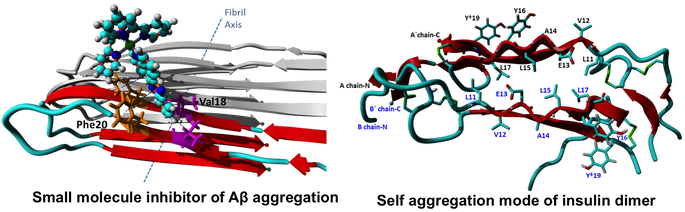
Studying protein self-oligomerization and design of small oligomerization inhibitors: Self-oligomerization of small bio-molecules lead to life threatening diseases, i.e. progression of Alzheimer’s disease caused by amyloid protein aggregation. Our research includes investigation of olimerization at the atomistic level. Upon gaining this knowledge we try to design small inhibitor molecules to interrupt protein-protein interactions utilizing techniques such as virtual screening and molecular docking simulations. These studies will enable the prodution of efficient aggregation inhibitors.
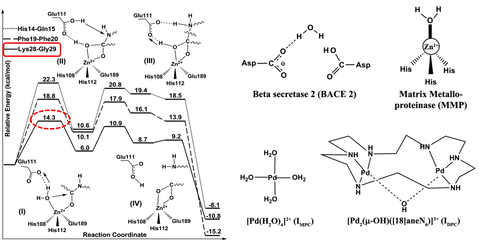
Investigation of enzyme mechanics and energetics: By using quantum mechanical (QM) calculations our group investigates the most plausible working mechanism of disease making proteins and enzymes. In addition, we also focus on increasing the activity of natural enzymes in tandem with experimentalists in the field of enzyme engineering.